An interdisciplinary team of mathematicians, engineers, physicists, and medical scientists have uncovered an unexpected link between pure mathematics and genetics, that reveals key insights into the structure of neutral mutations and the evolution of organisms.
Number theory, the study of the properties of positive integers, is perhaps the purest form of mathematics. At first sight, it may seem far too abstract to apply to the natural world. In fact, the influential American number theorist Leonard Dickson wrote, ‘Thank God that number theory is unsullied by any application.’ And yet, again and again, number theory finds unexpected applications in science and engineering, from leaf angles that (almost) universally follow the Fibonacci sequence, to modern encryption techniques based on factoring prime numbers. Now, researchers have demonstrated an unexpected link between number theory and evolutionary genetics.
Specifically, the team of researchers from Oxford, Harvard, Cambridge, GUST, MIT, Imperial, and the Alan Turing Institute, have discovered a deep connection between the sums-of-digits function from number theory and a key quantity in genetics, the phenotype mutational robustness. This quality is defined as the average probability that a point mutation does not change a phenotype (a characteristic of an organism).
Important implications
The discovery may have important implications for evolutionary genetics. Many genetic mutations are neutral, meaning that they can slowly accumulate over time without affecting the viability of the phenotype. These neutral mutations cause genome sequences to change at a steady rate over time. Because this rate is known, scientists can compare the percentage difference in the sequence between two organisms and infer when their latest common ancestor lived.
But the existence of these neutral mutations posed an important question: what fraction of mutations to a sequence are neutral? This property, called the phenotype mutational robustness, defines the average amount of mutations that can occur across all sequences without affecting the phenotype.
Maximum robustness
Professor Ard Louis from the University of Oxford, who led the study, said: ‘We have known for some time that many biological systems exhibit remarkably high phenotype robustness, without which evolution would not be possible. But we didn’t know what the absolute maximal robustness possible would be, or if there even was a maximum.’
It is precisely this question that the team has answered. They proved that the maximum robustness is proportional to the logarithm of the fraction of all possible sequences that map to a phenotype, with a correction which is given by the sums of digits function sk(n), defined as the sum of the digits of a natural number n in base k. For example, for n = 123 in base 10, the digit sum would be s10(123) = 1 + 2 + 3 = 6.
Tagaki function
Another surprise was that the maximum robustness also turns out to be related to the famous Tagaki function, a bizarre function that is continuous everywhere, but differentiable nowhere. This fractal function is also called the blancmange curve, because it looks like the French dessert.
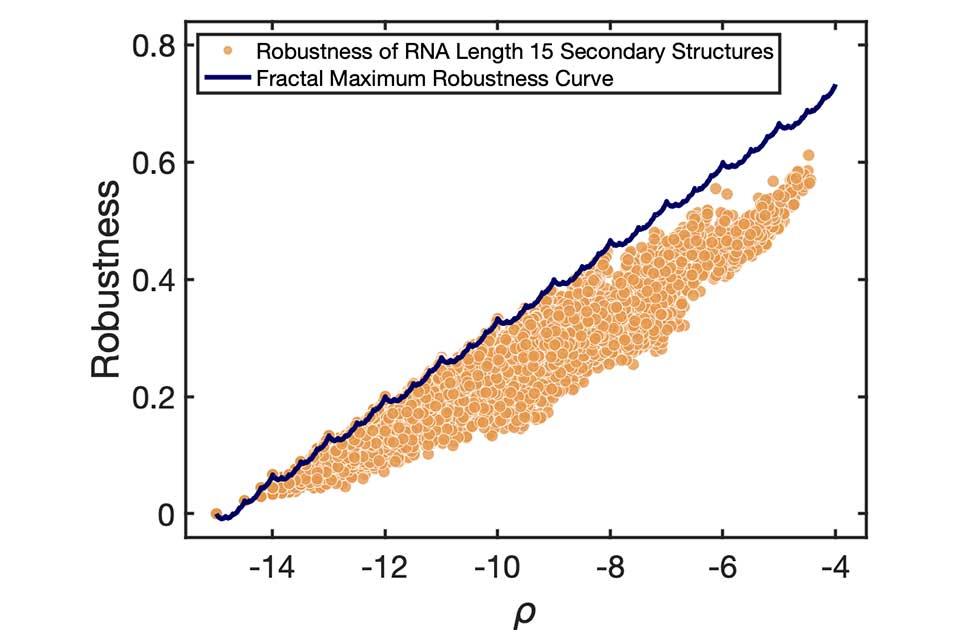
First author Dr. Vaibhav Mohanty, Harvard Medical School, adds: ‘What is most surprising is that we found clear evidence in the mapping from sequences to RNA secondary structures that nature in some cases achieves the exact maximum robustness bound. It’s as if biology knows about the fractal sums-of-digits function.’
Professor Ard Louis concludes: ‘The beauty of number theory lies not only in the abstract relationships it uncovers between integers, but also in the deep mathematical structures it illuminates in our natural world. We believe that many intriguing new links between number theory and genetics will be found in the future.’
Maximum mutational robustness in genotype–phenotype maps follows a self-similar blancmange-like curve, Vaibhav Mohanty et al, Journal of The Royal Society Interface, 26 July 2023