A Synthetic Peptide Mimic of λ-Cro shows Sequence-Specific Binding in Vitro and in Vivo
ACS Chemical Biology American Chemical Society (ACS) 7:6 (2012) 1084-1094
Peptide based Molecules as Protein-Protein Interaction Inhibitors: Tools for Chemical Genetics and Therapy
Current Chemical Biology Bentham Science Publishers 6:2 (2012) 145-163
A Genetic Network That Balances Two Outcomes Utilizes Asymmetric Recognition of Operator Sites
Biophysical Journal Elsevier 102:7 (2012) 1580-1589
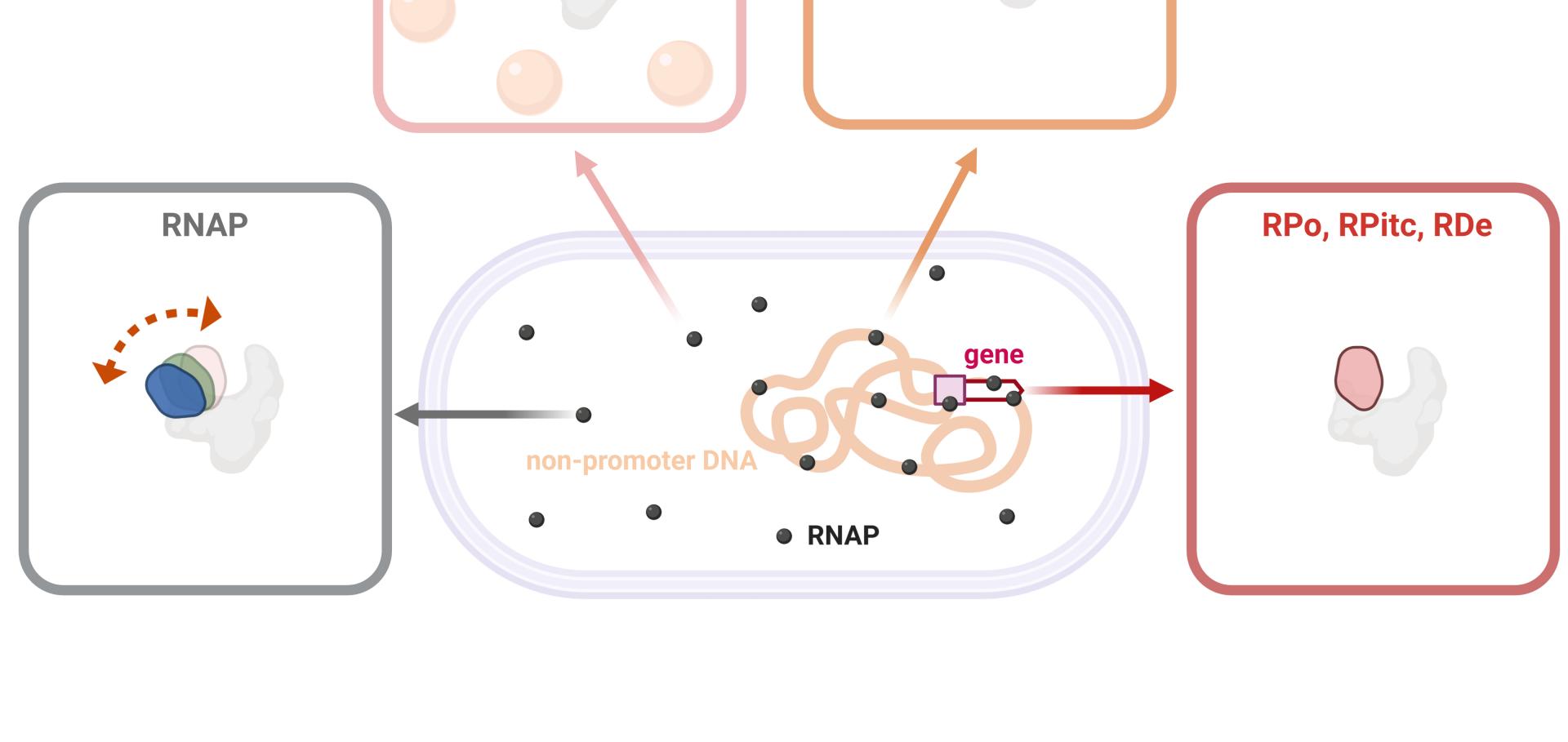
Recognition of different DNA sequences by a DNA‐binding protein alters protein dynamics differentially
FEBS Letters Wiley 586:3 (2012) 258-262